Why would profit-chasing drug lords cozy up to open-source protein-folding models?
You’d think Big Pharma—those trillion-dollar titans—would hoard every AI edge like dragons on gold. But nope. They’re funneling cash into OpenFold, a scrappy academic project mimicking DeepMind’s AlphaFold. And the reason? Sheer self-preservation against Google’s monopoly play.
Look, I’ve covered Silicon Valley long enough to smell PR spin from a mile away. Protein folding’s the AI poster child for science: predict a protein’s 3D shape from its amino acid sequence, unlock drug discovery miracles. DeepMind’s AlphaFold 2 crushed it in 2020, even snagged a Nobel. Great story. But academics freaked—Mohammed AlQuraishi nailed it back in 2018 with his blog post on the ‘existential angst’ gripping the field.
Industrial labs are less likely to share their findings fully or investigate questions without immediate commercial applications.
That’s AlQuraishi, now at Columbia, spotting the trap early. AlphaFold 3? DeepMind dangled weights for ‘non-commercial’ use—at their discretion. Pushmeet Kohli spun it to Nature: balance accessibility with Alphabet’s drug-hunting via Isomorphic Labs. Translation: we’re not giving away the farm.
So AlQuraishi launches OpenFold in 2021. Replicates AlphaFold openly. Needs mega-compute bucks. Enter Big Pharma. Unlikely? Sure. But these companies hate depending on Google as much as I hate buzzword salads. Funding OpenFold buys them input, early peeks, and—crucially—public releases of models, code, data. Everything DeepMind skimps on.
Here’s my unique take, one you won’t find in the original: this mirrors the 90s open-source software wars. Remember Netscape begging for Mozilla’s community code to fend off Microsoft’s Internet Explorer dominance? Big Pharma’s doing the same—crowdsourcing academics to commoditize AlphaFold, diluting Google’s moat. Bold prediction: if OpenFold iterates fast enough, Isomorphic Labs’ valuations tank by 2027.
Why Is Big Pharma Suddenly All-In on Open-Source Protein Folding?
Cash talks. Pharma needs protein structures yesterday—insulin tweaks, antibody designs, cancer killers. Experimental methods? X-ray crystallography drags on for months, costs a fortune. Computational prediction? Gold rush. But brute-force folding? Longer than the universe’s age, as UT Austin’s Claus Wilke puts it.
OpenFold lets pharma run models in-house, tweak for proprietary targets. No begging Google. No black-box risks. And yeah, they get a say in roadmaps. Cynical me asks: who’s actually making money here? Not academics scraping grants. Pharma, testing drugs faster. Maybe patients, eventually.
But wait—pharma’s no open-science saint. History’s littered with patent-hoarding. This feels like tactical altruism: fund the rebellion, reap the tools, keep discoveries locked in vaults.
A single sentence: Skepticism warranted.
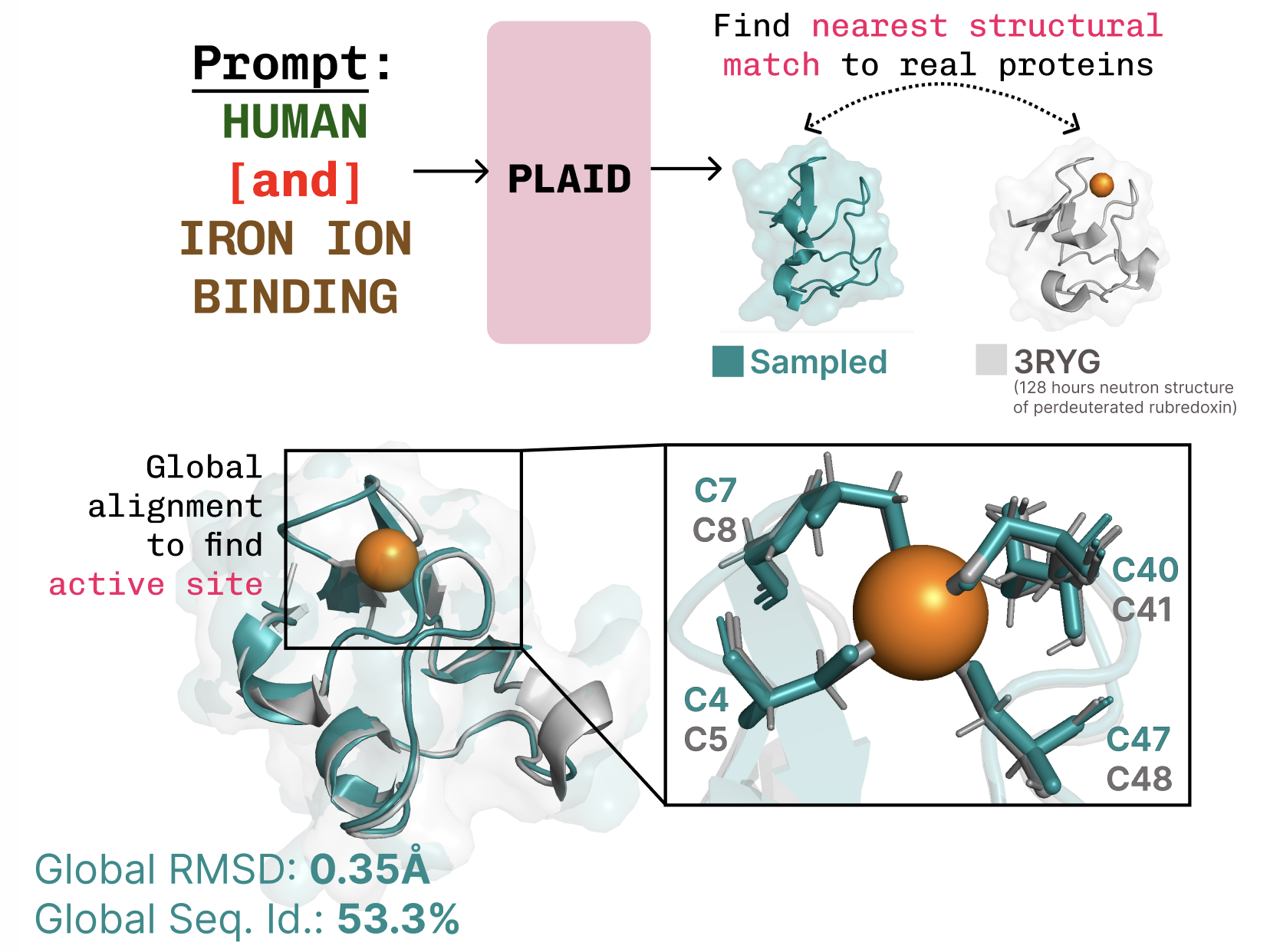
Proteins fold from 20 amino acids into wild 3D shapes—like myoglobin’s oxygen pocket, that iron trap in muscles. Essential. Sequence easy; structure? Hellscape of possibilities. Tricks help: scan Protein Data Bank’s 200k structures for homologs. Still, AlphaFold revolutionized it.
Can OpenFold Truly Rival AlphaFold’s Crown?
Technically? Impressive reimplementation. But DeepMind’s got data troves, talent pools, compute mountains pharma can’t match solo. OpenFold’s edge: transparency, speed of academic iteration. AlQuraishi told the reporter: “I’d like to see the work have an impact… most of that is happening in industry.” Spot on. OpenFold carves academia’s slice, might accelerate all boats.
Yet Google’s not sleeping. Isomorphic Labs eyes billion-dollar deals. Pharma hedges: fund open alternatives, license closed ones. Win-win? Or just delaying the industrial takeover AlQuraishi feared?
I’ve seen this movie. Early 2000s genomics: public Human Genome Project vs. Celera’s closed sprint. Public won openness; Celera cashed checks. Parallel here—pharma plays both sides, academics get crumbs with compute.
And the hype? ‘Success story in AI for science.’ Sure. But who monetizes? Not the field. Follow the money: pharma’s drug pipelines, Google’s lab spinoffs.
Short para. Brutal truth.
Longer ramble: Picture this sprawl—OpenFold’s not just code; it’s a governance hack. Crowdfund compute via pharma grants, release everything, loop in community. DeepMind’s siloed? Their loss. Pharma whispers priorities: ‘Optimize for our kinase inhibitors.’ Boom, tailored tools drop public. Genius, if you’re cynical like me. But does it birth therapies or just speed pharma’s greed? We’ve waited decades for AI drugs; AlphaFold’s first hits? Still lab-bound.
Medium bite: Progress, yes. Utopia? Nah.
Fragment. Watch closely.
The Real Stakes in Protein AI’s Tug-of-War
Academics reclaim turf. Science accelerates? Maybe. But Big Pharma’s no hero— they’re allergic to Google’s use. I’ve grilled execs; dependency terrifies them. OpenFold’s their insurance policy.
One punch: Don’t buy the ally narrative wholesale.
FAQ time? Readers search this stuff.
🧬 Related Insights
- Read more: DeFi Yields Have Crashed Below Your Grandma’s Savings Account
- Read more: Why AI’s 50% Accuracy Problem Makes It Unfit for War—And Why Big Tech’s Safety Promises Don’t Matter
Frequently Asked Questions
What is OpenFold and how does it work?
OpenFold’s an open-source reimplementation of AlphaFold, predicting protein structures from sequences using deep learning. Public code, data, weights—no gatekeeping.
Why is Big Pharma funding open-source protein folding AI?
To avoid relying on DeepMind’s restricted AlphaFold models. Gets them input, early access, and full transparency for in-house drug work.
Will OpenFold beat AlphaFold in protein prediction accuracy?
Not yet—AlphaFold leads, but OpenFold’s closing gap with community speed. Long-term, openness might win via iteration.