AlphaFold2 snagged the 2024 Nobel in Chemistry, capping a frenzy where AI cracked protein folding like never before. Markets buzzed: more predictors, faster structures from sequences. Billions poured into biotech on that promise.
PLAID upends it. This new model—PLAID, short for whatever acronym fits—learns diffusion in the latent space of folding giants like ESMFold. Result? Simultaneous generation of protein sequences (discrete strings) and full 3D all-atom structures (continuous coords). And here’s the kicker: it trains solely on sequences, tapping databases 2-4 orders of magnitude huger than scarce structural data.
PLAID is a multimodal generative model that simultaneously generates protein 1D sequence and 3D structure, by learning the latent space of protein folding models.
Expectations shattered. No more begging for cryo-EM shots or X-ray crystals—these are gold-standard structures, but experimental ones number maybe 200,000. Sequences? Hundreds of millions. PLAID drinks from that firehose, borrowing structural smarts from pretrained folders.
Why Bother Repurposing Folders for Generation?
Look, diffusion models have teased protein design before. But they choked on basics: backbone-only outputs, ignoring sidechains unless you already had the sequence. Multimodal mess. PLAID fixes it by sampling latents—frozen decoder spits out both.
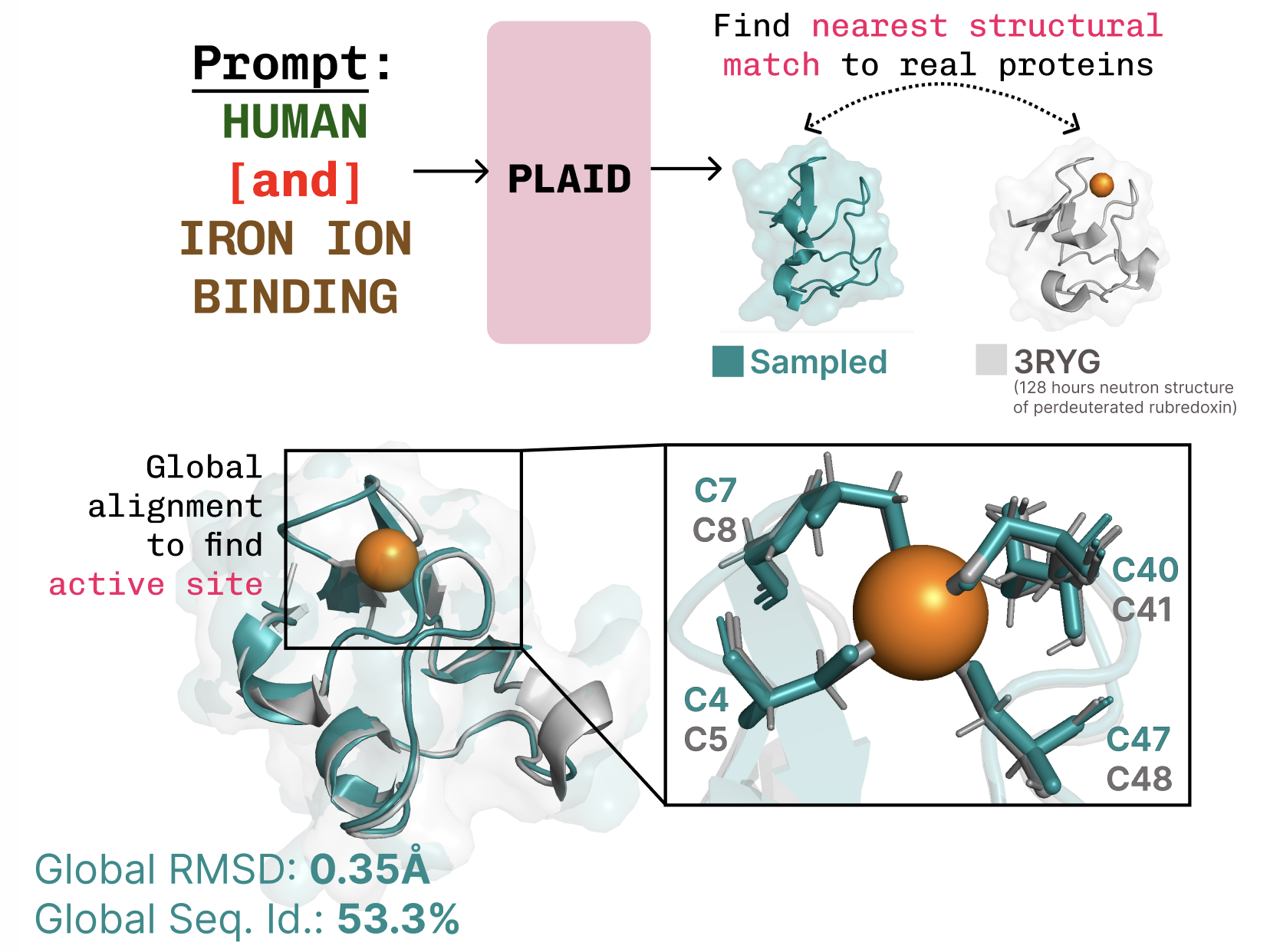
Organism tweaks, too. Human biologics can’t look alien; immune systems shred ‘em. PLAID takes prompts like “humanized” or “E. coli native,” compositional style akin to image gens (shoutout Liu et al., 2022). Function? It nails cysteine-iron coordination in metalloproteins, diverse sequences intact.
But. Is this hype? Folders like ESMFold embed sequences into latents packed with fold info. PLAID diffuses there, no structure labels needed during train. Inference decodes everything. Smart borrow, like robotics VLAs raiding internet-scale VLMs for priors.
ESMFold’s latents? Bloated, overactivated channels screaming for regularization. Enter CHEAP: their compression trick, hourglass-style, shrinks it down. Avoids the high-res image synth pitfalls.
One para punch: Scales beautifully.
Can PLAID Actually Design Real Drugs?
Drug discovery’s a slog—biology nailed, then solubility for pills over vials. How specify? Textual prompts, eventually. PLAID prototypes function + organism axes.
Data dynamics shift hard. Sequence DBs dwarf structures by 100-10,000x; cheaper too. Train generative prior on abundance, condition for utility. Pharma spends $2.6B per drug, 10-15 years. If PLAID halves that via controlled gens? Market cap boon—biotechs like Generate Biomedicines already value at $2B+ on similar bets.
Skepticism check: Generation’s easy; usefulness? Metrics show promise, but in vitro tests pending. Still, latent diffusion’s track record—from DALL-E to proteins—suggests momentum.
My take, data-driven: This echoes image diffusion’s pivot. Early classifiers (CLIP) got repurposed for gen via latents trained on billions. Unique insight—no one says it—PLAID’s the Stable Diffusion of proteins. Bold call: In 3 years, expect 10x cheaper custom biologics, licensing deals flooding from Big Pharma. But PR spin on “control”? Test it against immune evasion failures first.
And the method? Sequences -> ESMFold embed (frozen) -> diffusion in latent -> decode seq/struct. ❄️ frozen weights everywhere. Clean.
Visuals hint: Massive channel activations tamed by CHEAP. Top-3 outliers vs. medians—regularized bliss.
Why Does Sequence-Only Training Flip Protein AI?
Folding peaked; generation starves on data. Structures bottleneck—expensive, sparse. Sequences flow free.
PLAID sidesteps. Learns distribution from seqs alone, folds implicitly via pretrained latents. Broader coverage, too—exotic organisms, functions underrepresented in PDB.
Market angle: Biotech VC hit $20B in 2023, AI slice growing 50% YoY. PLAID-like tools? Accelerant. But watch: Discrete-continuous coupling tricky; diffusion stumbles on modes. Their multimodal fix—impressive, unproven at scale.
From prediction to design. Nobel marked endgame? Nah. PLAID signals therapeutics goldrush.
Short burst: Game on.
Deeper: Imagine interfaces—prompt “soluble humanized enzyme for Fe catalysis, tablet-friendly.” Outputs diverse candidates, structures ready. Iterate fast.
Limits linger. No full text control yet. Organism prompts basic. But proof-of-concept lands.
Historical parallel—GANs started toy-ish, morphed markets. PLAID? Same trajectory, bioflavor.
🧬 Related Insights
- Read more: o3’s 10x RL Compute Gambit: The Real State of LLM Reasoning Reinforcement
- Read more: Claude Code Agents in Parallel: Worktrees End the Waiting Game
Frequently Asked Questions
What is PLAID in protein AI?
PLAID’s a generative model that creates new protein sequences and 3D structures by diffusing in the latent space of models like ESMFold, trained only on abundant sequence data.
How does PLAID generate proteins without structures?
It borrows structural knowledge from pretrained folding models’ frozen latents—sample there, decode both seq and struct at inference.
Will PLAID speed up drug discovery?
Potentially huge: Larger training data, prompt control for functions/organisms. Could slash costs, but real-world validation needed amid immune and solubility hurdles.